Open data lakehouse for biology
Context and memory for datasets and models at scale, across infrastructure, and open-source.
Query, trace & validate with a lineage-native lakehouse that supports bio-formats, registries & ontologies.
From the creators of



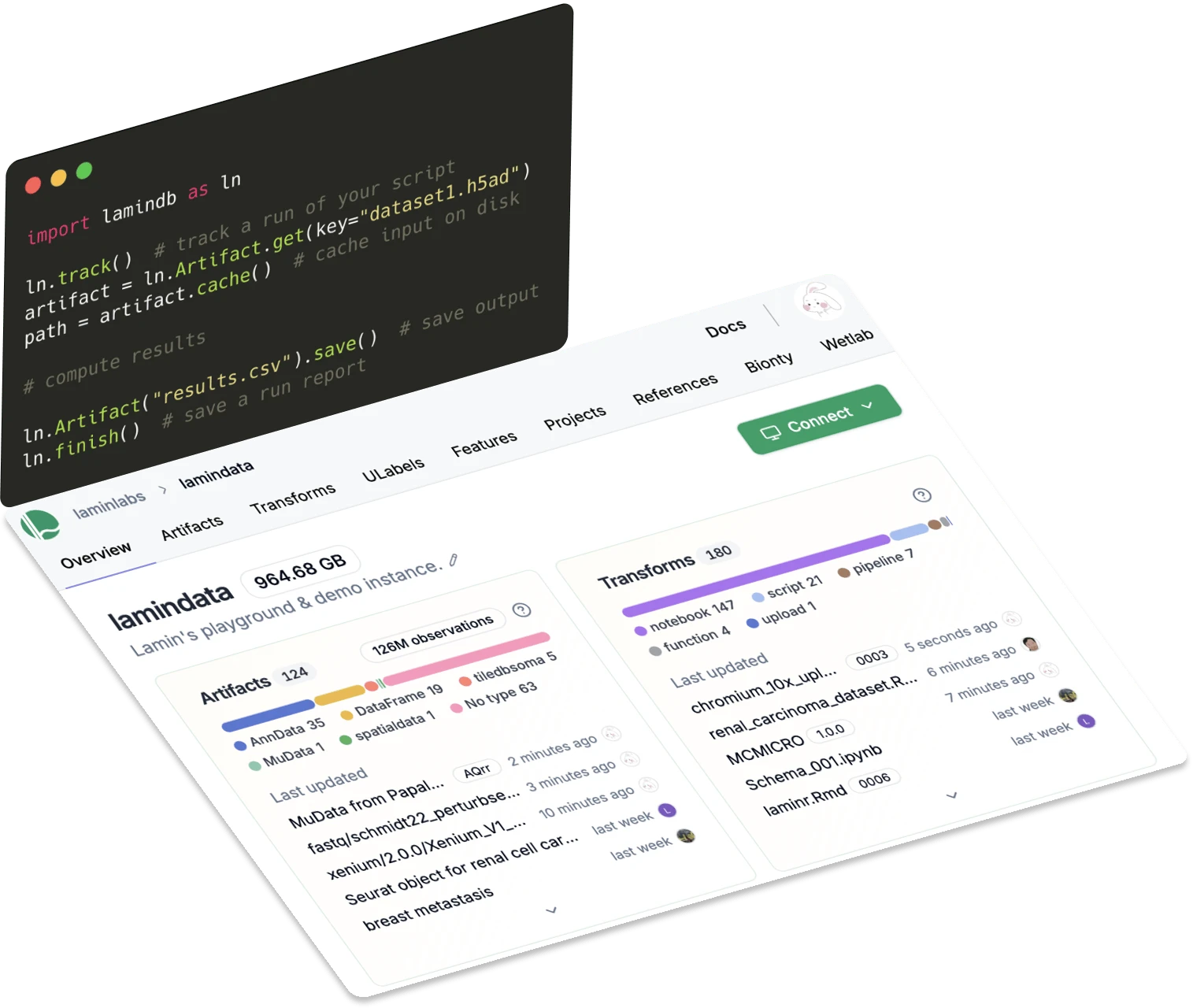
Lineage
Trace data, code & reports
Know where data came from and what it's used for. Track data lineage with a single line of code.
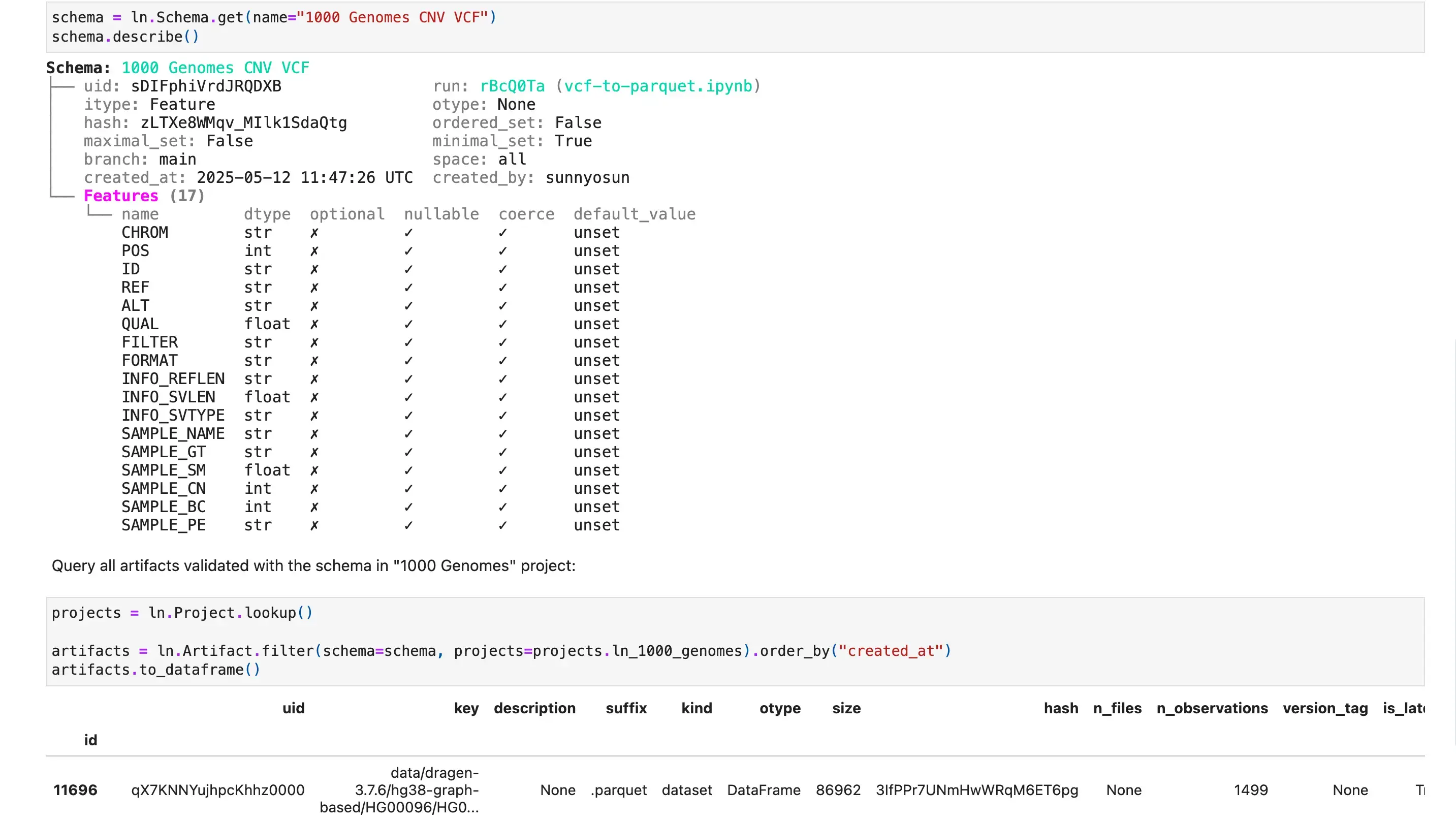
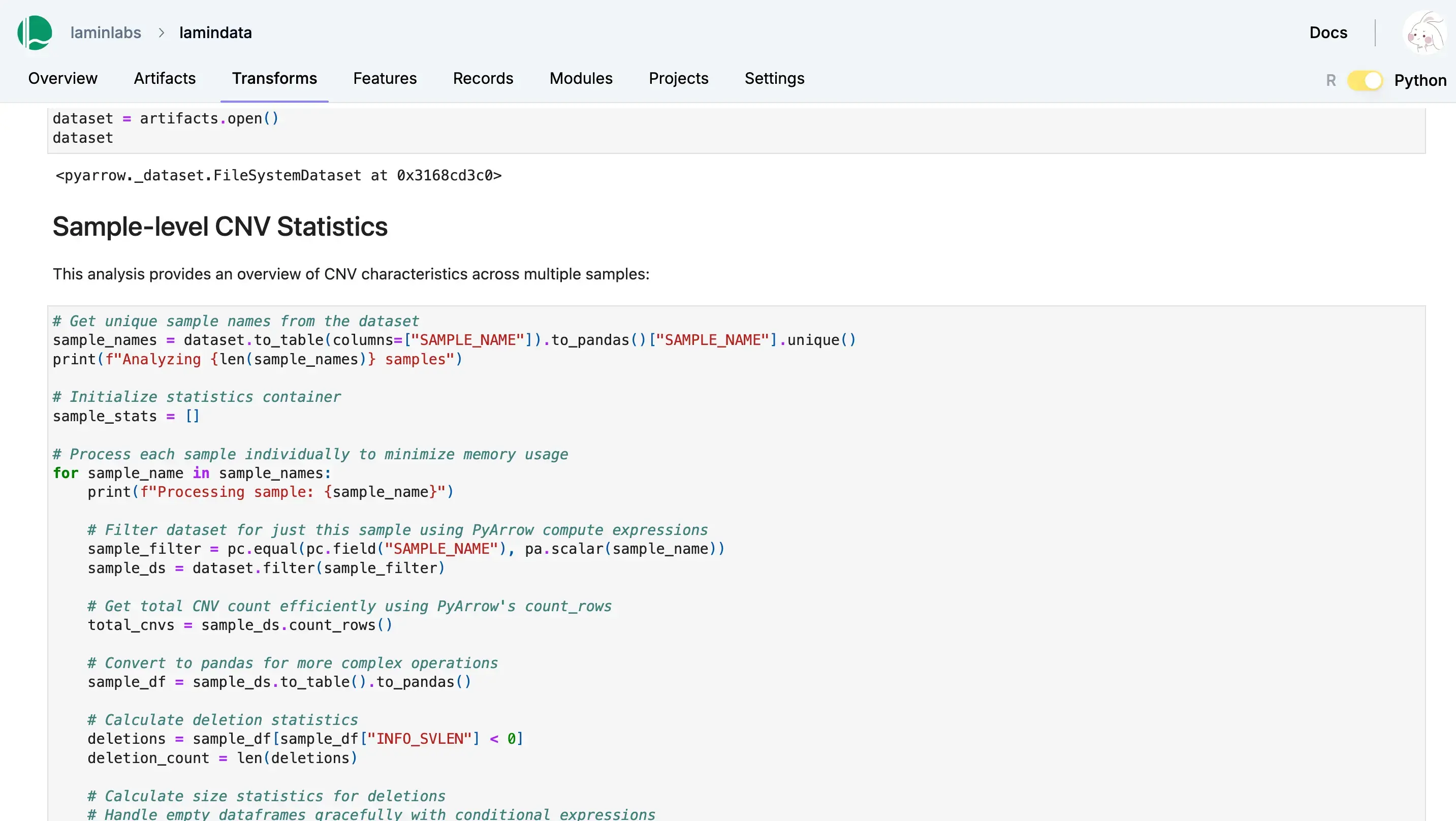
Lakehouse + Bio-formats
Query datasets at scale
Query and batch-load datasets with lakehouse support for a wide range of table & array formats. Manage their features & schemas as metadata in Postgres or SQLite.
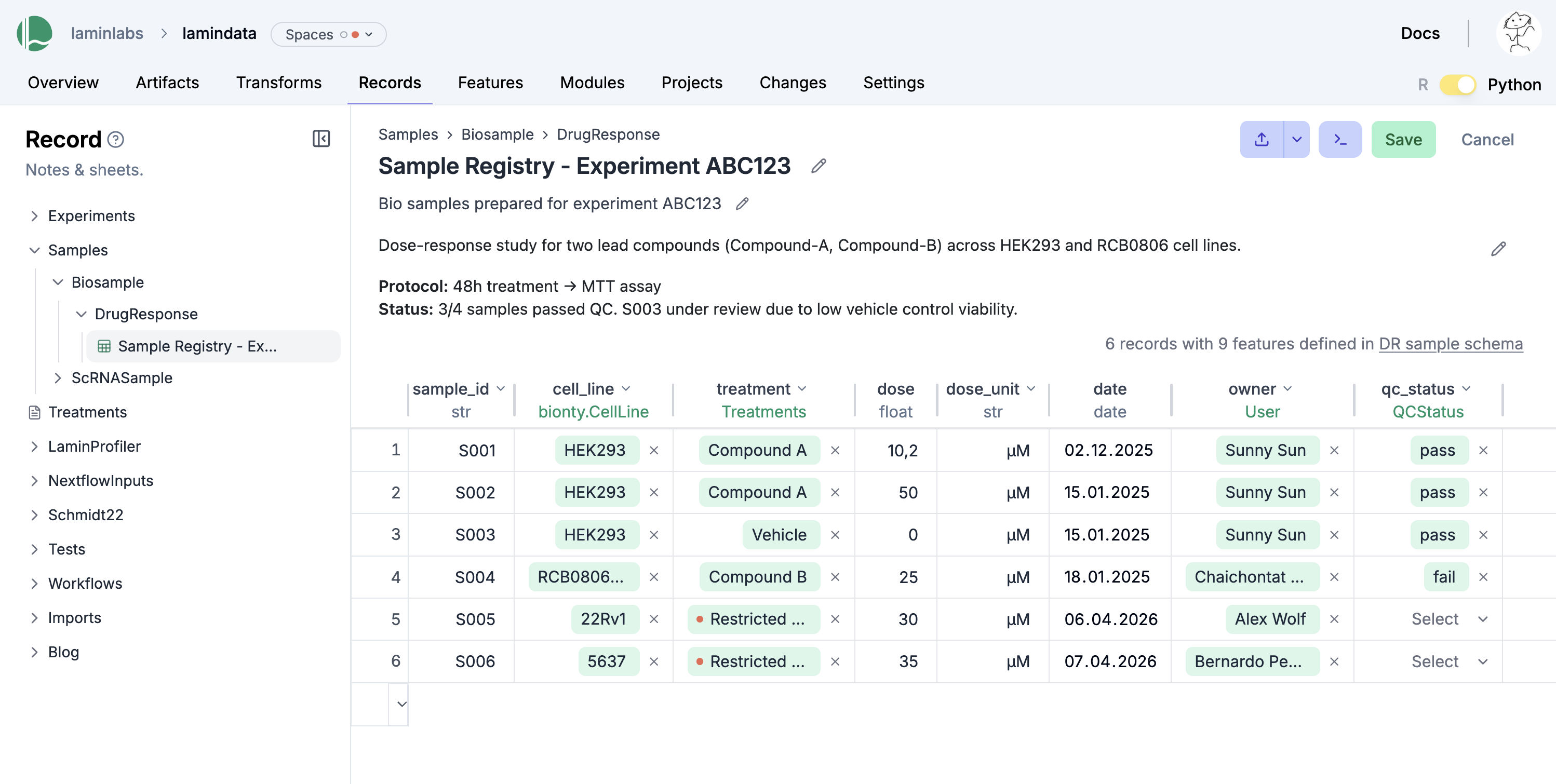
Registries + Sheets (LIMS)
Unify metadata and datasets
Manage metadata in relational sheets in sync with datasets in storage. Use a single Python/R class with built-in ontologies, project & change management.
Integrity
Validate & annotate datasets
Use schemas to enforce consistency across your data assets. Annotate datasets with a single line of code.
Zero lock-in
Administer with ease while staying in control
Manage fine-grained permissions for humans & agents with SaaS-like simplicity directly at the database and storage level. Do not give up admin control on AWS, GCP, or in your own infrastructure.
Context
Build your organization's long-term memory
As team & agents work, data, models & reports get mapped into the lakehouse — building recursively queryable memory & training data that compounds over time.