CELLxGENE: scRNA-seq#
CZ CELLxGENE hosts the globally largest standardized collection of scRNA-seq datasets.
LaminDB makes it easy to query the CELLxGENE data and integrate it with in-house data of any kind (omics, phenotypes, pdfs, notebooks, ML models, …).
You can use the CELLxGENE data in three ways:
In the current guide, you’ll see how to query metadata and data based on
AnnDataobjects.If you want to use these in your own LaminDB instance, see the transfer guide.
If you’d like to leverage the TileDB-SOMA API for the data subset of CELLxGENE Census, see the Census guide.
If you are interested in building similar data assets in-house:
See the scRNA guide for how to create a growing versioned queryable scRNA-seq dataset.
See the Annotate for validating, curating and registering your own AnnData objects.
Reach out if you are interested in a full zero-copy clone of
laminlabs/cellxgeneto accelerate building your in-house LaminDB instances.
Setup#
Load the public LaminDB instance that mirrors cellxgene on the CLI:
!lamin load laminlabs/cellxgene
💡 connected lamindb: laminlabs/cellxgene
import lamindb as ln
import bionty as bt
💡 connected lamindb: laminlabs/cellxgene
❗ Full backed capabilities are not available for this version of anndata, please install anndata>=0.9.1.
Query & understand metadata#
Auto-complete metadata#
You can create look-up objects for any registry in LaminDB, including basic biological entities and things like users or storage locations.
Let’s use auto-complete to look up cell types:
Show me a screenshot
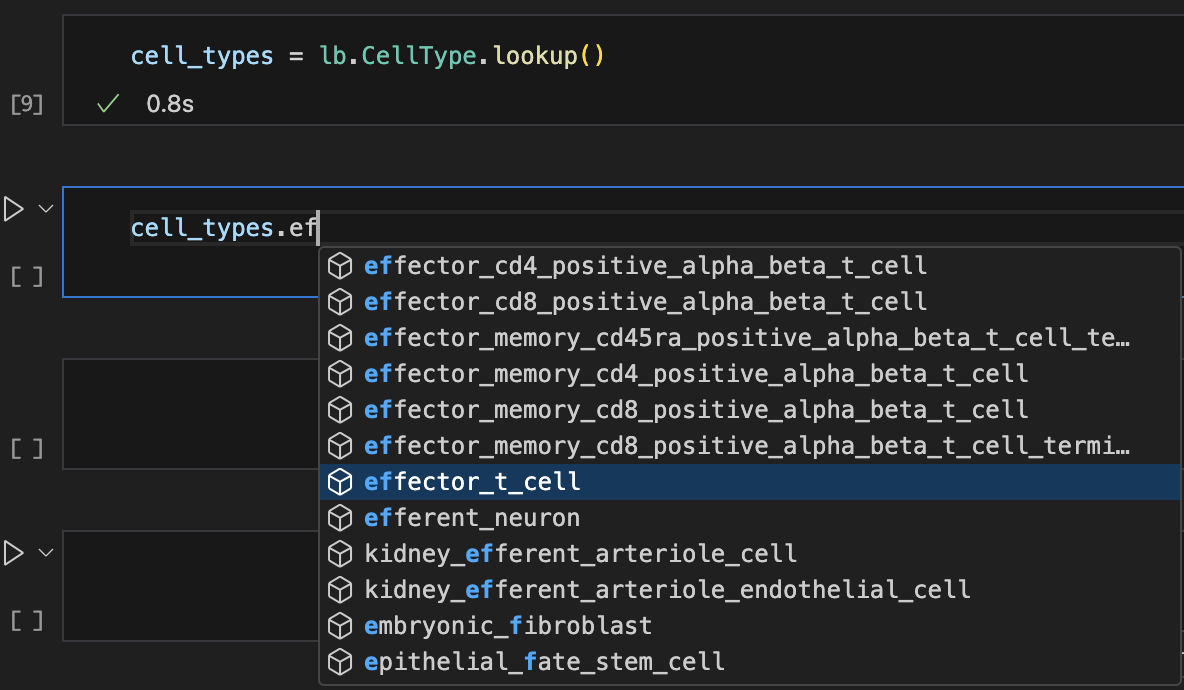
cell_types = bt.CellType.lookup()
cell_types.effector_t_cell
CellType(uid='3nfZTVV4', name='effector T cell', ontology_id='CL:0000911', synonyms='effector T-cell|effector T-lymphocyte|effector T lymphocyte', description='A Differentiated T Cell With Ability To Traffic To Peripheral Tissues And Is Capable Of Mounting A Specific Immune Response.', updated_at=2023-11-28 22:30:57 UTC, public_source_id=48, created_by_id=1)
You can also arbitrarily chain filters and create lookups from them:
organisms = bt.Organism.lookup()
experimental_factors = bt.ExperimentalFactor.lookup() # labels for experimental factors
tissues = bt.Tissue.lookup() # tissue labels
suspension_types = ln.ULabel.filter(name="is_suspension_type").one().children.lookup() # suspension types
Search & filter metadata#
We can use search & filters for metadata:
bt.CellType.search("effector T cell")
Show code cell output
| uid | synonyms | score | |
|---|---|---|---|
| name | |||
| effector T cell | 3nfZTVV4 | effector T-cell|effector T-lymphocyte|effector... | 100.0 |
| ectodermal cell | 2rFEBLPn | ectoderm cell | 71.4 |
| helper T cell | 43cBCa7s | helper T-lymphocyte|T-helper cell|helper T lym... | 71.4 |
| memory T cell | 1oa5G2Mq | memory T-cell|memory T lymphocyte|memory T-lym... | 71.4 |
| sensory receptor cell | 6GkjRSiR | receptor cell | 71.4 |
| excretory cell | 5teqLp2U | 69.0 | |
| secretory cell | 4eEkKmdU | 69.0 | |
| neurectodermal cell | 1eJqfkLq | neurectoderm cell | 68.8 |
| pro-T cell | 4twkhtZN | pro-T lymphocyte|progenitor T cell | 68.8 |
| regulatory T cell | 6IELBVIu | regulatory T lymphocyte|Treg|regulatory T-lymp... | 68.8 |
| Kupffer cell | 5fdXwyLs | hepatic macrophage|macrophagocytus stellatus|l... | 66.7 |
| chemoreceptor cell | 6wMTbaYL | 66.7 | |
| follicular B cell | 2EhFTUoZ | Fo B cell|follicular B lymphocyte|follicular B... | 66.7 |
bt.CellType.search("CD8-positive cytokine effector T cell")
Show code cell output
| uid | synonyms | score | |
|---|---|---|---|
| name | |||
| CD8-positive, alpha-beta cytokine secreting effector T cell | 6JD5JCZC | CD8-positive, alpha-beta cytokine secreting ef... | 77.1 |
| CD4-positive helper T cell | 531hEapj | CD4-positive T-helper cell|CD4-positive helper... | 69.8 |
| CD8-positive, alpha-beta T cell | 6IC9NGJE | CD8-positive, alpha-beta T-cell|CD8-positive, ... | 67.6 |
| CD8-positive, alpha-beta cytotoxic T cell | Mv6woHvO | CD8-positive, alpha-beta cytotoxic T-cell|CD8-... | 66.7 |
| CD8-positive, alpha-beta memory T cell | 7MuNkhO9 | CD8-positive, alpha-beta memory T lymphocyte|C... | 66.7 |
| CD1c-positive myeloid dendritic cell | 5fo7nTlc | 65.8 | |
| Tc1 cell | 4AQr9CRo | Tc1 T lymphocyte|Tc1 T-cell|Tc1 T cell|T-cytot... | 65.5 |
| CD141-positive myeloid dendritic cell | CAwwMhIV | 64.9 | |
| CD4-positive, alpha-beta T cell | 4PSMdO3I | CD4-positive, alpha-beta T lymphocyte|CD4-posi... | 64.7 |
| CD4-positive, alpha-beta cytotoxic T cell | 5zRXDnpu | CD4-positive, alpha-beta cytotoxic T-cell|CD4-... | 64.1 |
And use a uid to filter exactly one metadata record:
effector_t_cell = bt.CellType.filter(uid="3nfZTVV4").one()
effector_t_cell
CellType(uid='3nfZTVV4', name='effector T cell', ontology_id='CL:0000911', synonyms='effector T-cell|effector T-lymphocyte|effector T lymphocyte', description='A Differentiated T Cell With Ability To Traffic To Peripheral Tissues And Is Capable Of Mounting A Specific Immune Response.', updated_at=2023-11-28 22:30:57 UTC, public_source_id=48, created_by_id=1)
Understand ontologies#
View the related ontology terms:
effector_t_cell.view_parents(distance=2, with_children=True)
Or access them programmatically:
effector_t_cell.children.df()
| uid | name | ontology_id | abbr | synonyms | description | public_source_id | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|
| id | ||||||||||
| 931 | 2VQirdSp | effector CD8-positive, alpha-beta T cell | CL:0001050 | None | effector CD8-positive, alpha-beta T lymphocyte... | A Cd8-Positive, Alpha-Beta T Cell With The Phe... | 48 | 2023-11-28 22:27:55.565976+00:00 | 2023-11-28 22:27:55.565981+00:00 | 1 |
| 1088 | 490Xhb24 | effector CD4-positive, alpha-beta T cell | CL:0001044 | None | effector CD4-positive, alpha-beta T lymphocyte... | A Cd4-Positive, Alpha-Beta T Cell With The Phe... | 48 | 2023-11-28 22:27:55.569828+00:00 | 2023-11-28 22:27:55.569832+00:00 | 1 |
| 1229 | 69TEBGqb | exhausted T cell | CL:0011025 | None | Tex cell|An effector T cell that displays impa... | None | 48 | 2023-11-28 22:27:55.572880+00:00 | 2023-11-28 22:27:55.572884+00:00 | 1 |
| 1309 | 5s4gCMdn | cytotoxic T cell | CL:0000910 | None | cytotoxic T lymphocyte|cytotoxic T-lymphocyte|... | A Mature T Cell That Differentiated And Acquir... | 48 | 2023-11-28 22:27:55.575440+00:00 | 2023-11-28 22:27:55.575444+00:00 | 1 |
| 1331 | 43cBCa7s | helper T cell | CL:0000912 | None | helper T-lymphocyte|T-helper cell|helper T lym... | A Effector T Cell That Provides Help In The Fo... | 48 | 2023-11-28 22:27:55.575949+00:00 | 2023-11-28 22:27:55.575955+00:00 | 1 |
Query artifacts#
Unlike in the SOMA guide, here, we’ll query sets of .h5ad files, which correspond to AnnData objects.
To access them, we query the Collection record that links the latest LTS set of .h5ad artifacts:
collection = ln.Collection.filter(name="cellxgene-census", version="2023-12-15").one()
collection
Collection(uid='dMyEX3NTfKOEYXyMu591', name='cellxgene-census', version='2023-12-15', hash='0NB32iVKG5ttaW5XILvG', visibility=1, updated_at=2024-01-30 09:09:49 UTC, transform_id=19, run_id=24, created_by_id=1)
You can get all linked artifacts as a dataframe - there are >1000 h5ad files in cellxgene-census version 2023-12-15.
collection.artifacts.count()
1113
collection.artifacts.df().head() # not tracking run & transform because read-only instance
Show code cell output
| uid | storage_id | key | suffix | accessor | description | version | size | hash | hash_type | n_objects | n_observations | transform_id | run_id | visibility | key_is_virtual | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||||||||
| 2825 | OoktqBIu8jCoGOJlaQPo | 2 | cell-census/2023-12-15/h5ads/fc0ceb80-d2d9-47c... | .h5ad | AnnData | Sst Chodl - DLPFC: Seattle Alzheimer's Disease... | 2023-12-15 | 73375840 | DqV7FraZIIP_l2DJuvHk_g-9 | md5-n | None | 1877 | 16 | 22 | 1 | False | 2024-01-11 09:13:25.387366+00:00 | 2024-01-24 07:18:54.197599+00:00 | 1 |
| 2031 | n33nFE2kXSNzNhIAtS3S | 2 | cell-census/2023-12-15/h5ads/44c83972-e5d2-485... | .h5ad | AnnData | L5 IT - DLPFC: Seattle Alzheimer's Disease Atl... | 2023-12-15 | 4605202922 | ztuPyGXWH_OyCq1OyPlNkw-549 | md5-n | None | 104106 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.820851+00:00 | 2024-01-24 07:19:02.027481+00:00 | 1 |
| 1813 | mtoOxeGG0Rg3NPH1AlwD | 2 | cell-census/2023-12-15/h5ads/100c6145-7b0e-4ba... | .h5ad | AnnData | Microglia-PVM - DLPFC: Seattle Alzheimer's Dis... | 2023-12-15 | 634716733 | -B96CrmiOANuzE3xU78WsQ-76 | md5-n | None | 42486 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.307694+00:00 | 2024-01-24 07:19:04.190720+00:00 | 1 |
| 1804 | V0tqrgE1z1NY2eUUKKQE | 2 | cell-census/2023-12-15/h5ads/0ed60482-a34f-426... | .h5ad | AnnData | Lamp5 - DLPFC: Seattle Alzheimer's Disease Atl... | 2023-12-15 | 1580667477 | xRTDQGA4iOC4r8sSgz53vQ-189 | md5-n | None | 55968 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.282158+00:00 | 2024-01-24 07:19:04.646675+00:00 | 1 |
| 2532 | dEP0dZ8UxLgwnkLjHssX | 2 | cell-census/2023-12-15/h5ads/bd65a70f-b274-413... | .h5ad | AnnData | Single-cell sequencing links multiregional imm... | 2023-12-15 | 1204103287 | 5hUwdflh_erDK-U2bEzfvw-144 | md5-n | None | 167283 | 16 | 22 | 1 | False | 2024-01-11 09:13:24.792407+00:00 | 2024-01-29 07:49:54.125887+00:00 | 1 |
You can query across artifacts by arbitrary metadata combinations, for instance:
query = collection.artifacts.filter(
organism=organisms.human,
cell_types__in=[cell_types.dendritic_cell, cell_types.neutrophil],
tissues=tissues.kidney,
ulabels=suspension_types.cell,
experimental_factors=experimental_factors.ln_10x_3_v2,
)
query = query.order_by("size") # order by size
query.df().head() # convert to DataFrame
Show code cell output
| uid | storage_id | key | suffix | accessor | description | version | size | hash | hash_type | n_objects | n_observations | transform_id | run_id | visibility | key_is_virtual | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||||||||
| 1880 | WwmBIhBNLTlRcSoBpatT | 2 | cell-census/2023-12-15/h5ads/20d87640-4be8-487... | .h5ad | AnnData | Mature kidney dataset: immune | 2023-12-15 | 44647761 | hSLF-GPhLXaC2tVIOJEdXA-6 | md5-n | None | 7803 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.448150+00:00 | 2024-01-29 07:46:33.152678+00:00 | 1 |
| 1880 | WwmBIhBNLTlRcSoBpatT | 2 | cell-census/2023-12-15/h5ads/20d87640-4be8-487... | .h5ad | AnnData | Mature kidney dataset: immune | 2023-12-15 | 44647761 | hSLF-GPhLXaC2tVIOJEdXA-6 | md5-n | None | 7803 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.448150+00:00 | 2024-01-29 07:46:33.152678+00:00 | 1 |
| 1930 | gHlQ5Muwu3G9pvFC7egT | 2 | cell-census/2023-12-15/h5ads/2d31c0ca-0233-41c... | .h5ad | AnnData | Fetal kidney dataset: immune | 2023-12-15 | 64056560 | jENeQIq0JdoHl5PyfY-sjA-8 | md5-n | None | 6847 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.544310+00:00 | 2024-01-29 07:46:37.205210+00:00 | 1 |
| 2405 | P4Oai3OLGAzRwoicaxCB | 2 | cell-census/2023-12-15/h5ads/9ea768a2-87ab-46b... | .h5ad | AnnData | Mature kidney dataset: full | 2023-12-15 | 192484358 | yghldeu2bOC5jtvnqZH8Og-23 | md5-n | None | 40268 | 16 | 22 | 1 | False | 2024-01-11 09:13:24.526987+00:00 | 2024-01-29 07:49:11.905786+00:00 | 1 |
| 2405 | P4Oai3OLGAzRwoicaxCB | 2 | cell-census/2023-12-15/h5ads/9ea768a2-87ab-46b... | .h5ad | AnnData | Mature kidney dataset: full | 2023-12-15 | 192484358 | yghldeu2bOC5jtvnqZH8Og-23 | md5-n | None | 40268 | 16 | 22 | 1 | False | 2024-01-11 09:13:24.526987+00:00 | 2024-01-29 07:49:11.905786+00:00 | 1 |
Query arrays#
Each artifact stores an array in form of an annotated data matrix, an AnnData object.
Let’s look at the first array in the artifact query and show metadata using .describe():
artifact = query.first()
artifact.describe()
Show code cell output
Artifact(uid='WwmBIhBNLTlRcSoBpatT', key='cell-census/2023-12-15/h5ads/20d87640-4be8-487f-93d4-dce38378d00f.h5ad', suffix='.h5ad', accessor='AnnData', description='Mature kidney dataset: immune', version='2023-12-15', size=44647761, hash='hSLF-GPhLXaC2tVIOJEdXA-6', hash_type='md5-n', n_observations=7803, visibility=1, key_is_virtual=False, updated_at=2024-01-29 07:46:33 UTC)
Provenance:
📎 storage: Storage(uid='oIYGbD74', root='s3://cellxgene-data-public', type='s3', region='us-west-2')
📎 transform: Transform(uid='V4AGIdOJcOgj6K79', name='Census release 2023-12-15 (LTS)', key='cencus-release-2023-12-15-LTS', version='0', type='notebook')
📎 run: Run(uid='UAAiLAi0BrLvlKnsuvP3', started_at=2024-01-29 07:27:05 UTC, is_consecutive=False)
📎 created_by: User(uid='kmvZDIX9', handle='sunnyosun', name='Sunny Sun')
📎 input_of (core.Run): ['2024-01-30 09:07:36 UTC']
Features:
var: FeatureSet(uid='MLFo2ZBXvibkOyBR9UOR', n=32922, type='number', registry='bionty.Gene')
'PKIG', 'CXCL6', 'KIAA0586', 'C5orf24', 'MAP4K3', 'ZDHHC7', 'OR2J2', 'PPWD1', 'CHRM4', 'SPRYD3', 'RASSF5', 'PAX3', 'TNFRSF14-AS1', 'RPS10P7', 'TMEM88B', 'WBP1', 'MED28-DT', 'MCCD1', 'RIBC1', 'RAB31', ...
obs: FeatureSet(uid='zAQ6WnmIMDLslhfgdIOt', name='obs metadata', n=11, type='category', registry='core.Feature')
🔗 tissue (5, bionty.Tissue): 'kidney', 'kidney blood vessel', 'renal medulla', 'renal pelvis', 'cortex of kidney'
🔗 tissue_type (0, core.ULabel):
🔗 assay (1, bionty.ExperimentalFactor): '10x 3' v2'
🔗 cell_type (12, bionty.CellType): 'classical monocyte', 'non-classical monocyte', 'neutrophil', 'mature NK T cell', 'kidney resident macrophage', 'CD8-positive, alpha-beta T cell', 'dendritic cell', 'B cell', 'natural killer cell', 'mast cell', ...
🔗 development_stage (12, bionty.DevelopmentalStage): '49-year-old human stage', '64-year-old human stage', '67-year-old human stage', '72-year-old human stage', '53-year-old human stage', '12-year-old human stage', '19-month-old human stage', '2-year-old human stage', '63-year-old human stage', '4-year-old human stage', ...
🔗 disease (1, bionty.Disease): 'normal'
🔗 donor_id (13, core.ULabel): 'pRCC', 'TTx', 'TxK4', 'RCC1', 'Wilms3', 'TxK3', 'TxK2', 'TxK1', 'Wilms1', 'VHL', ...
🔗 self_reported_ethnicity (1, bionty.Ethnicity): 'unknown'
🔗 sex (2, bionty.Phenotype): 'male', 'female'
🔗 suspension_type (1, core.ULabel): 'cell'
🔗 organism (1, bionty.Organism): 'human'
Labels:
📎 organism (1, bionty.Organism): 'human'
📎 tissues (5, bionty.Tissue): 'kidney', 'kidney blood vessel', 'renal medulla', 'renal pelvis', 'cortex of kidney'
📎 cell_types (12, bionty.CellType): 'classical monocyte', 'non-classical monocyte', 'neutrophil', 'mature NK T cell', 'kidney resident macrophage', 'CD8-positive, alpha-beta T cell', 'dendritic cell', 'B cell', 'natural killer cell', 'mast cell', ...
📎 diseases (1, bionty.Disease): 'normal'
📎 phenotypes (2, bionty.Phenotype): 'male', 'female'
📎 experimental_factors (1, bionty.ExperimentalFactor): '10x 3' v2'
📎 developmental_stages (12, bionty.DevelopmentalStage): '49-year-old human stage', '64-year-old human stage', '67-year-old human stage', '72-year-old human stage', '53-year-old human stage', '12-year-old human stage', '19-month-old human stage', '2-year-old human stage', '63-year-old human stage', '4-year-old human stage', ...
📎 ethnicities (1, bionty.Ethnicity): 'unknown'
📎 ulabels (14, core.ULabel): 'pRCC', 'TTx', 'TxK4', 'RCC1', 'Wilms3', 'TxK3', 'TxK2', 'TxK1', 'Wilms1', 'VHL', ...
More ways of accessing metadata
Access just features:
artifact.features
Or get labels given a feature:
artifact.labels.get(features.tissue).df()
artifact.labels.get(features.collection).one()
If you want to query a slice of the array data, you have two options:
Cache & load the entire array into memory via
artifact.load() -> AnnData(caches the h5ad on disk, so that you only download once)Stream the array from the cloud using a cloud-backed accessor
artifact.backed() -> AnnDataAccessor
Both options will run much faster if you run them close to the data (AWS S3 on the US West Coast, consider logging into hosted compute there).
Cache & load:
adata = artifact.load()
adata
Show code cell output
AnnData object with n_obs × n_vars = 7803 × 32922
obs: 'donor_id', 'donor_age', 'self_reported_ethnicity_ontology_term_id', 'organism_ontology_term_id', 'sample_uuid', 'tissue_ontology_term_id', 'development_stage_ontology_term_id', 'suspension_uuid', 'suspension_type', 'library_uuid', 'assay_ontology_term_id', 'mapped_reference_annotation', 'is_primary_data', 'cell_type_ontology_term_id', 'author_cell_type', 'disease_ontology_term_id', 'reported_diseases', 'sex_ontology_term_id', 'compartment', 'Experiment', 'Project', 'cell_type', 'assay', 'disease', 'organism', 'sex', 'tissue', 'self_reported_ethnicity', 'development_stage'
var: 'feature_is_filtered', 'feature_name', 'feature_reference', 'feature_biotype'
uns: 'default_embedding', 'schema_version', 'title'
obsm: 'X_umap'
Now we have an AnnData object, which stores observation annotations matching our artifact-level query in the .obs slot, and we can re-use almost the same query on the array-level.
See the array-level query
adata_slice = adata[
adata.obs.cell_type.isin(
[cell_types.dendritic_cell.name, cell_types.neutrophil.name]
)
& (adata.obs.tissue == tissues.kidney.name)
& (adata.obs.suspension_type == suspension_types.cell.name)
& (adata.obs.assay == experimental_factors.ln_10x_3_v2.name)
]
adata_slice
See the artifact-level query for comparison
query = collection.artifacts.filter(
organism=organisms.human,
cell_types__in=[cell_types.dendritic_cell, cell_types.neutrophil],
tissues=tissues.kidney,
ulabels=suspension_types.cell,
experimental_factors=experimental_factors.ln_10x_3_v2,
)
AnnData uses pandas to manage metadata and the syntax differs slightly. However, the same metadata records are used.
Stream:
adata_backed = artifact.backed()
adata_backed
Show code cell output
AnnDataAccessor object with n_obs × n_vars = 7803 × 32922
constructed for the AnnData object 20d87640-4be8-487f-93d4-dce38378d00f.h5ad
obs: ['Experiment', 'Project', '_index', 'assay', 'assay_ontology_term_id', 'author_cell_type', 'cell_type', 'cell_type_ontology_term_id', 'compartment', 'development_stage', 'development_stage_ontology_term_id', 'disease', 'disease_ontology_term_id', 'donor_age', 'donor_id', 'is_primary_data', 'library_uuid', 'mapped_reference_annotation', 'organism', 'organism_ontology_term_id', 'reported_diseases', 'sample_uuid', 'self_reported_ethnicity', 'self_reported_ethnicity_ontology_term_id', 'sex', 'sex_ontology_term_id', 'suspension_type', 'suspension_uuid', 'tissue', 'tissue_ontology_term_id']
obsm: ['X_umap']
raw: ['X', 'var', 'varm']
uns: ['default_embedding', 'schema_version', 'title']
var: ['_index', 'feature_biotype', 'feature_is_filtered', 'feature_name', 'feature_reference']
We now have an AnnDataAccessor object, which behaves much like an AnnData, and the query looks the same.
See the query
adata_backed_slice = adata_backed[
adata_backed.obs.cell_type.isin(
[cell_types.dendritic_cell.name, cell_types.neutrophil.name]
)
& (adata_backed.obs.tissue == tissues.kidney.name)
& (adata_backed.obs.suspension_type == suspension_types.cell.name)
& (adata_backed.obs.assay == experimental_factors.ln_10x_3_v2.name)
]
adata_backed_slice.to_memory()
Train an ML model#
You can directly train an ML models on the entire collection.
Exploring data by collection#
Alternatively,
you can search a file on the LaminHub UI and fetch it through:
ln.Artifact.get(uid)or query for a collection you found on CZ CELLxGENE Discover
Let’s search the collections from CELLxGENE within the 2023-12-15 release:
ln.Collection.filter(version="2023-12-15").search("immune human kidney", limit=10)
| uid | score | |
|---|---|---|
| name | ||
| Spatiotemporal immune zonation of the human kidney | kqiPjpzpK9H9rdtnV67f | 55.1 |
| The integrated Human Lung Cell Atlas | FaJmPleTV3HjPBTdFyOZ | 43.6 |
| Asian Immune Diversity Atlas (AIDA) | ZzUntpjOt8v7Awqdkpja | 41.5 |
| Single cell derived mRNA signals across human kidney tumors | Yed6da6CsPXaGmLQDTBi | 41.0 |
| Live Human Microglia Single-cell RNA-seq | olY10cghAPIz2oGrQvRC | 40.7 |
| Human Brain Cell Atlas v1.0 | kDJ9Xb8d11d93LAHZLTC | 39.1 |
| Azimuth meta-analysis of human scRNA-seq datasets | RoCAhVTi0ao0p5yX5GZE | 38.2 |
| Distinct microbial and immune niches of the human colon | VVsweEynenmLLY85Bvvx | 37.8 |
| Human developing neocortex by area | 6tT3kRYI2c6slEpvtOWS | 37.7 |
| Mapping the developing human immune system across organs | veK7yRfThpdptF4vPT3l | 37.3 |
Let’s get the record of the top hit collection:
collection = ln.Collection.filter(uid="kqiPjpzpK9H9rdtnV67f").one()
collection
Collection(uid='kqiPjpzpK9H9rdtnV67f', name='Spatiotemporal immune zonation of the human kidney', description='10.1126/science.aat5031', version='2023-12-15', hash='4wGcXeeqsjVdbRdU7ZuJ', reference='120e86b4-1195-48c5-845b-b98054105eec', reference_type='CELLxGENE Collection ID', visibility=1, updated_at=2024-01-29 07:54:33 UTC, transform_id=17, run_id=22, created_by_id=1)
We see it’s a Science paper and we could find more information using the DOI or CELLxGENE collection id.
Check different versions of this collection:
collection.versions.df()
| uid | name | description | version | hash | reference | reference_type | transform_id | run_id | artifact_id | visibility | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | ||||||||||||||
| 17 | kqiPjpzpK9H9rdtnHWas | Spatiotemporal immune zonation of the human ki... | 10.1126/science.aat5031 | 2023-07-25 | w_VZE7n841ktaA9FjdLh | 120e86b4-1195-48c5-845b-b98054105eec | CELLxGENE Collection ID | NaN | NaN | None | 1 | 2024-01-08 12:01:20.121086+00:00 | 2024-01-08 12:01:20.121095+00:00 | 1 |
| 365 | kqiPjpzpK9H9rdtnV67f | Spatiotemporal immune zonation of the human ki... | 10.1126/science.aat5031 | 2023-12-15 | 4wGcXeeqsjVdbRdU7ZuJ | 120e86b4-1195-48c5-845b-b98054105eec | CELLxGENE Collection ID | 17.0 | 22.0 | None | 1 | 2024-01-11 13:41:06.531224+00:00 | 2024-01-29 07:54:33.854515+00:00 | 1 |
Each collection has at least one Artifact file associated to it. Let’s get the associated artifacts:
collection.artifacts.df()
| uid | storage_id | key | suffix | accessor | description | version | size | hash | hash_type | n_objects | n_observations | transform_id | run_id | visibility | key_is_virtual | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||||||||
| 1778 | b2x19Eg28GGSNnXW1hAD | 2 | cell-census/2023-12-15/h5ads/08073b32-d389-41f... | .h5ad | AnnData | Fetal kidney dataset: nephron | 2023-12-15 | 159545411 | _JE59jFHDrOn0hj4i1yXSQ-20 | md5-n | None | 10790 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.214114+00:00 | 2024-01-29 07:46:06.497662+00:00 | 1 |
| 1880 | WwmBIhBNLTlRcSoBpatT | 2 | cell-census/2023-12-15/h5ads/20d87640-4be8-487... | .h5ad | AnnData | Mature kidney dataset: immune | 2023-12-15 | 44647761 | hSLF-GPhLXaC2tVIOJEdXA-6 | md5-n | None | 7803 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.448150+00:00 | 2024-01-29 07:46:33.152678+00:00 | 1 |
| 1930 | gHlQ5Muwu3G9pvFC7egT | 2 | cell-census/2023-12-15/h5ads/2d31c0ca-0233-41c... | .h5ad | AnnData | Fetal kidney dataset: immune | 2023-12-15 | 64056560 | jENeQIq0JdoHl5PyfY-sjA-8 | md5-n | None | 6847 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.544310+00:00 | 2024-01-29 07:46:37.205210+00:00 | 1 |
| 1944 | USUgRVwrCMquHiImhk5D | 2 | cell-census/2023-12-15/h5ads/2fc9c59f-3cfd-48d... | .h5ad | AnnData | Mature kidney dataset: non PT parenchyma | 2023-12-15 | 39294782 | 3l5iNnBmPFbYfR3-THYWNQ-5 | md5-n | None | 4620 | 16 | 22 | 1 | False | 2024-01-11 09:13:23.568572+00:00 | 2024-01-29 07:46:52.173865+00:00 | 1 |
| 2405 | P4Oai3OLGAzRwoicaxCB | 2 | cell-census/2023-12-15/h5ads/9ea768a2-87ab-46b... | .h5ad | AnnData | Mature kidney dataset: full | 2023-12-15 | 192484358 | yghldeu2bOC5jtvnqZH8Og-23 | md5-n | None | 40268 | 16 | 22 | 1 | False | 2024-01-11 09:13:24.526987+00:00 | 2024-01-29 07:49:11.905786+00:00 | 1 |
| 2570 | 6mnZ3SeQFhffr3wTdZZb | 2 | cell-census/2023-12-15/h5ads/c52de62a-058d-4d7... | .h5ad | AnnData | Fetal kidney dataset: stroma | 2023-12-15 | 109942751 | s24Q5-FNUNQPLZw9BuwOVg-14 | md5-n | None | 8345 | 16 | 22 | 1 | False | 2024-01-11 09:13:24.870820+00:00 | 2024-01-29 07:50:01.866851+00:00 | 1 |
| 2652 | 11HQaMeIUaOwyHoOWVvA | 2 | cell-census/2023-12-15/h5ads/d7dcfd8f-2ee7-438... | .h5ad | AnnData | Fetal kidney dataset: full | 2023-12-15 | 341214674 | 2mnG5TiEpj0Wr5L19TTFRw-41 | md5-n | None | 27197 | 16 | 22 | 1 | False | 2024-01-11 09:13:25.042157+00:00 | 2024-01-29 07:50:28.610568+00:00 | 1 |
